Parrish Lab, University of Washington (2010-Present)
2022
Drosophila epidermal cells are intrinsically mechanosensitive and drive nociceptive behavioral outputs
Jiro Yoshino, Sonali S. Mali, Claire R. Williams, Takeshi Morita, Chloe E. Emerson, Christopher J. Arp, Sophie E. Miller, Chang Yin, Lydia Thé, Chikayo Hemmi, Mana Motoyoshi, Kenichi Ishii, Kazuo Emoto, Diana M. Bautista, and Jay Z. Parrish (2022). bioRxiv, DOI: 10.1101/2022.10.07.511265
2021
Transparent touch: Insights from model systems on epidermal control of somatosensory innervation
Chang Yin, Eric Peterman, Jeffrey P Rasmussen, Jay Z Parrish (2021). Frontiers in Cellular Neuroscience, DOI: 10.1101/2022.10.07.511265
2020
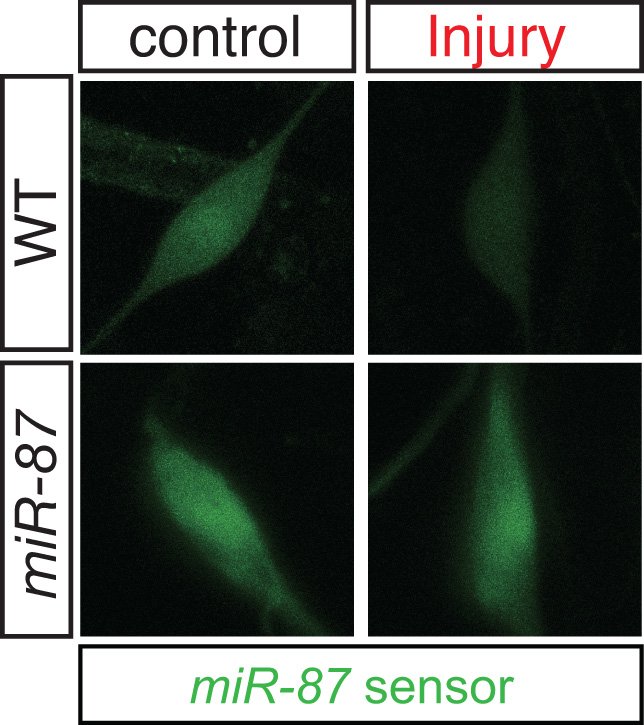
Drosophila miR-87 promotes dendrite regeneration by targeting the transcriptional repressor Tramtrack69
Yasuko Kitatani, Akane Tezuka, Eri Hasegawa, Satoyoshi Yanagi, Kazuya Togashi, Masato Tsuji, Shu Kondo, Jay Z Parrish, Kazuo Emoto (2020). PLoS Genetics 16(8)e1008942 DOI: 10.1371/journal.pgen.1008942.
Conserved Tao kinase activity regulates dendrite arborization, cytoskeletal dynamics, and sensory function in Drosophila
Chun Hu, Alexandros K Kanellopoulos, Melanie Richter, Meike Petersen, Anja Konietzny, Federico M Tenedini, Nina Hoyer, Lin Cheng, Carole LC Poon, Kieran F Harvey, Sabine Windhorst, Jay Z Parrish, Marina Mikhaylova, Claudia Bagni, Froylan Calderon de Anda, Peter Soba (2020) Journal of Neuroscience DOI: 10.1523/JNEUROSCI.1846-19.2020
Think Globally, Act Locally: Scaling the Growth of Motor Neurons
Jiro Yoshino, Kazuo Emoto, and Jay Z. Parrish (2020). Developmental Cell 54(1) 5-6. DOI: 10.1016/j.devcel.2020.06.015
2019
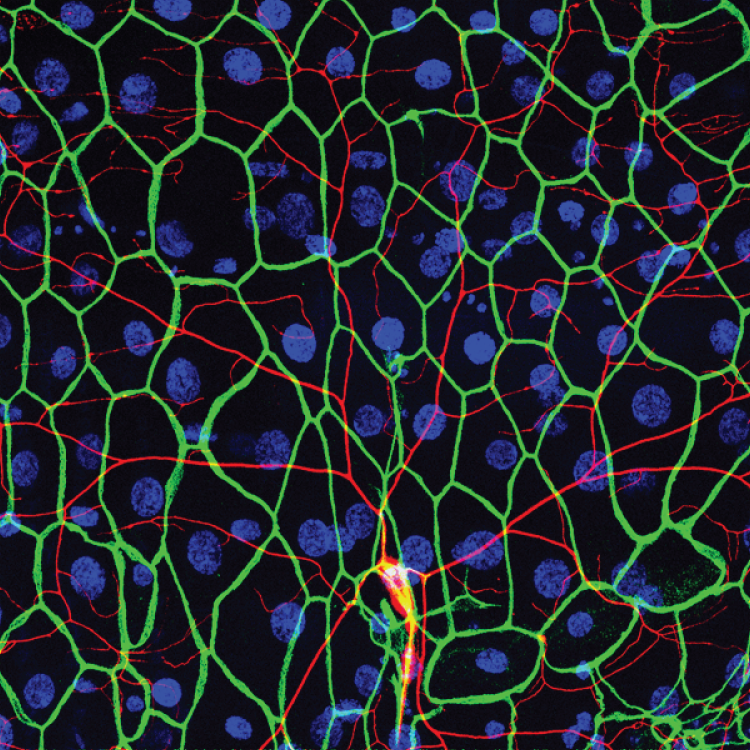
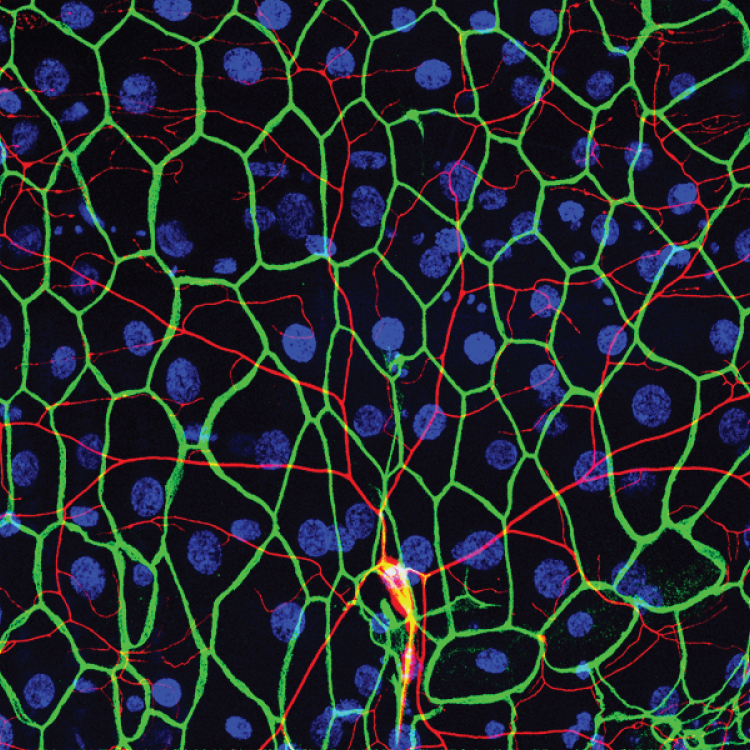
A conserved morphogenetic mechanism for epidermal ensheathment of nociceptive sensory neurites
Nan Jiang, Jeffrey P Rasmussen, Joshua A Clanton, Marci Rosenberg, Kory P Luedke, Mark R Cronan, Edward Parker, Hyeon-Jin Kim, Joshua C Vaughan, Alvaro Sagasti, Jay Z. Parrish (2019). eLife 8:e42455 DOI: 10.7554/eLife.42455.
2018
Empirical assessment of the impact of sample number and read depth on RNA-Seq analysis workflow performance
Alyssa Baccarella, Claire R. Williams, Jay Z. Parrish, Charles C. Kim (2018). BMC Bioinformatics 19:423, https://doi.org/10.1186/s12859-018-2445-2.
Microtubule acetylation is required for mechanosensation in Drosophila
Connie Yan, Fei Wang, Yun Peng, Claire R. Williams, Brian Jenkins, Jill Wildonger, John C. Tuthill, Yang Xiang, Stephen L. Rogers†, Jay Z. Parrish† (2018). Cell Reports 25:1051-1065.e6.
Ret and Substrate-Derived TGF-beta Maverick Regulate Space-Filling Dendrite Growth in Drosophila Sensory Neurons
Nina Hoyer, Philip Zielke, Chun Hu, Meike Petersen, Kathrin Sauter, Robin Scharrenberg, Yun Peng, Charles C. Kim, Chun Han, Jay Z. Parrish, and Peter Soba. (2018) Cell Reports 24:2261-2275 (Cover article)
Superresolution imaging of Drosophila tissue using expansion microscopy
Nan Jiang, Hyeon-Jin Kim, Tyler J. Chozinski, Jorge E. Azpurua, Benjamin A. Eaton, Joshua C. Vaughan†, and Jay Z. Parrish† (2018) Molecular Biology of the Cell 29:1413-1421 †correspondence (Cover Article)
Modulation of host learning in Aedes aegypti mosquitos
Clement Vinauger, Chloe Lahondère, Gabriella H. Wolff, Lauren T. Locke, LT, Jessica E. Liaw, JE, Jay Z. Parrish, Omar S. Akbari, Michael H. Dickinson, Jeffrey A. Riffell (2018) Current Biology 28: 333-344.
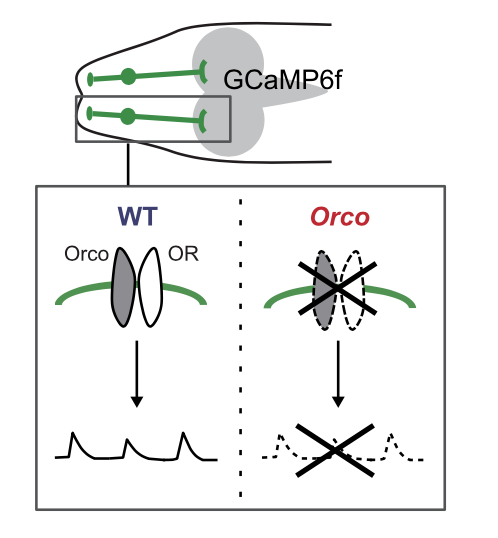
Prior activity of olfactory receptor neurons is required for proper sensory processing and behavior in Drosophila larvae
Nao Utashiro, Claire R. Williams, Jay Z. Parrish, and Kazuo Emoto (2018) Scientific Reports 8: 8580.
2017
TrpA1 activation in peripheral sensory neurons underlies the ionic basis of pain hypersensitivity in response to vinca alkaloids
Nina Boiko, Geraldo Medrano, Elizabeth Mantano, Nan Jiang, Claire R. Williams, Charles C. Kim, Jay Z. Parrish, Kenneth M. Hargreaves, James D. Stockand, and Benjamin A. Eaton (2017) PLoS One 12(10):e0186888, PMID: 29084244
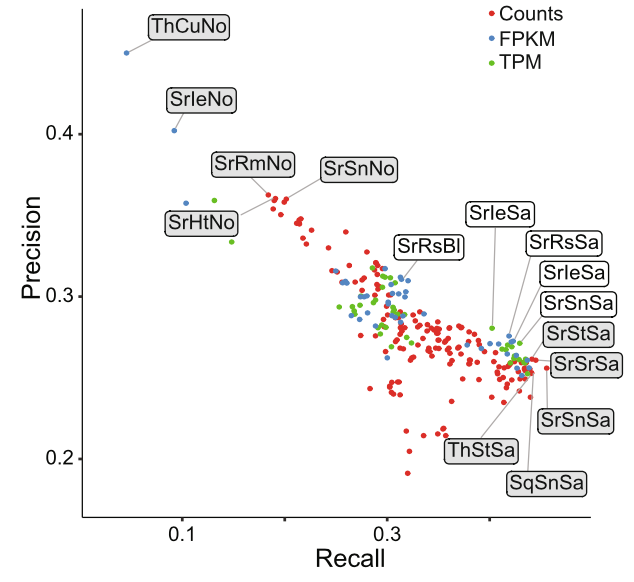
Empirical Assessment of Analysis Workflows for differential expression analysis of human samples using RNA-Seq
Claire R. Williams, Alyssa Baccarella, Jay Z. Parrish, Charles C. Kim (2017). BMC Bioinformatics 18: 38. PMID: 28095772
See additional coverage in RNA-seq blog here.
2016
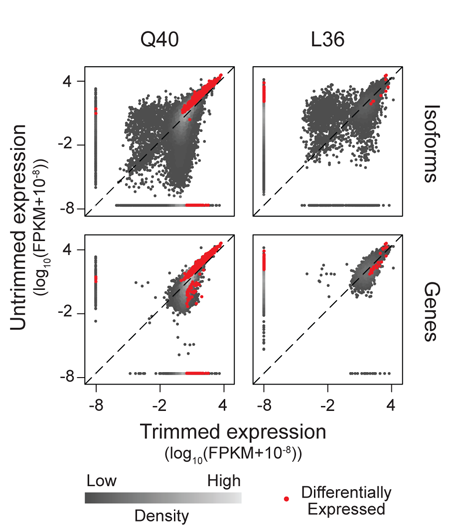
Trimming of Sequence Reads Alters RNA-Seq Gene Expression Estimates
Claire R. Williams, Alyssa Baccarella, Jay Z. Parrish†, Charles C. Kim† (2016). BMC Bioinformatics 17: 103. PMID: 26911985 †authors for correspondence
See additional coverage in RNA-seq blog here.
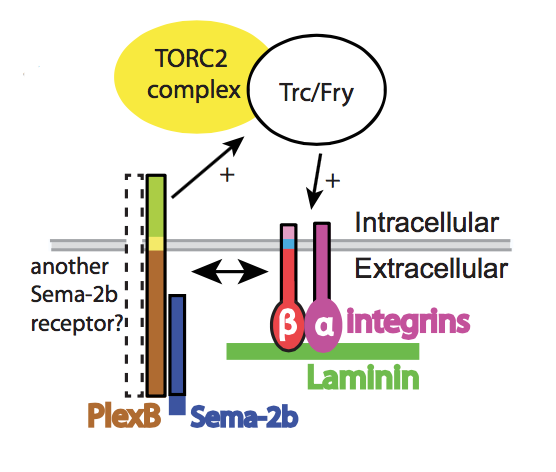
Epidermis-derived Semaphorin Promotes Dendrite Self-Avoidance by Regulating Dendrite-Substrate Adhesion in Drosophila Sensory Neurons
Shan Meltzer, Smita Yadav, Jiae Lee, Peter Soba, Susan H. Younger, Peng Jin, Wei Zhang, Jay Parrish, Lily Yeh Jan, Yuh Nung Jan (2016). Neuron 89: 741-755. PMID: 26853303.
See also Sci Signaling Editors Choice: Epidermal signals confine dendrites
Functions of the SLC36 transporter Pathetic in growth control.
Lin, WY, Williams, CR, Yan, C, and Parrish, JZ (2016). Fly doi: 10.1080/19336934.2015.1129089
http://www.ncbi.nlm.nih.gov/pubmed/26735916
Tiling and Mosaic Spacing of Dendrites
Jay Z. Parrish (2016). Chapter in Dendrites: Development and Disease. Emoto, Hoogenraad, Huang, and Wang (Eds.), Springer Press.
2015
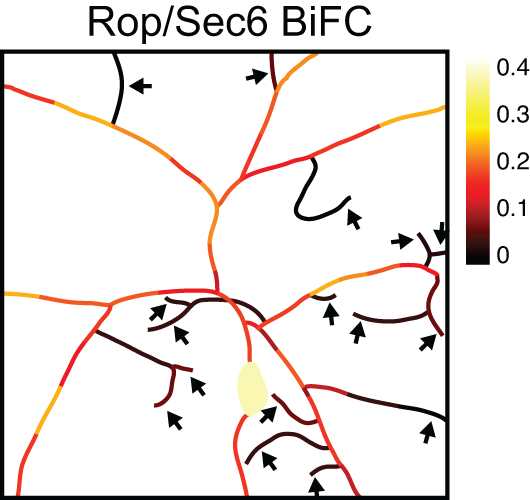
Regulation of dendrite growth and maintenance by exocytosis.
Yun Peng*, Jiae Lee*, Kimberly Rowland, Yuhui Wen, Hope Hua, Nicole Carlson, Shweta Lavania, Jay Z. Parrish†, Michael D. Kim† (2015). Journal of Cell Science 128: 4279-4292. PMCID: PMC4712815 *authors contributed equally, †authors for correspondence
See also JCS highlights: Rop controls dendrite growth
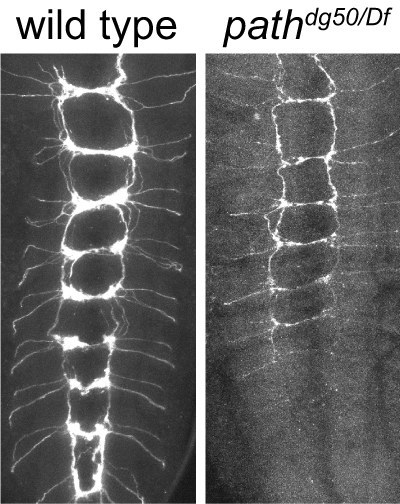
The SLC36 transporter pathetic is required for extreme growth in Drosophila neurons
Lin, W.Y., Williams, C., Luedke, K.A., Yan, C., Ahn, J., Morrison, N., Bloomsburg, S., Duncan, K., Kim, C.K., and Parrish, J.Z. (2015). Genes & Development 29:1120-1135. PMCID: PMC4470281.
http://www.ncbi.nlm.nih.gov/pubmed/26063572
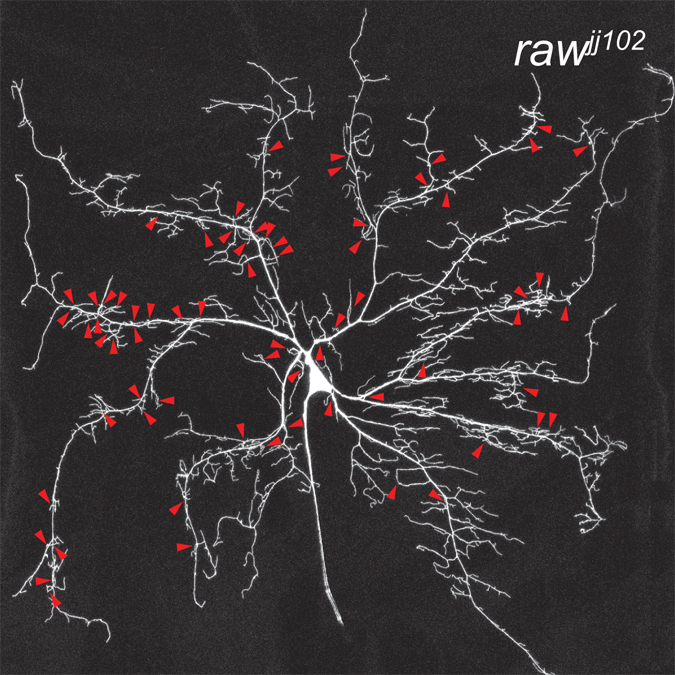
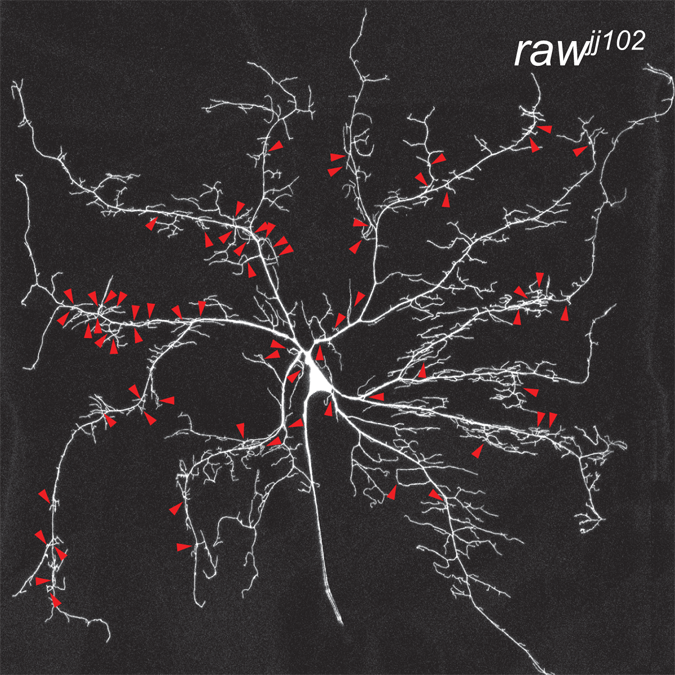
Coordinate control of terminal dendrite patterning and dynamics by the novel membrane protein Raw
Lee, J., Lin, W., and Parrish, J.Z. (2015). Development 142:1-12. PMCID: PMC25480915.
http://www.ncbi.nlm.nih.gov/pubmed/25480915
2014
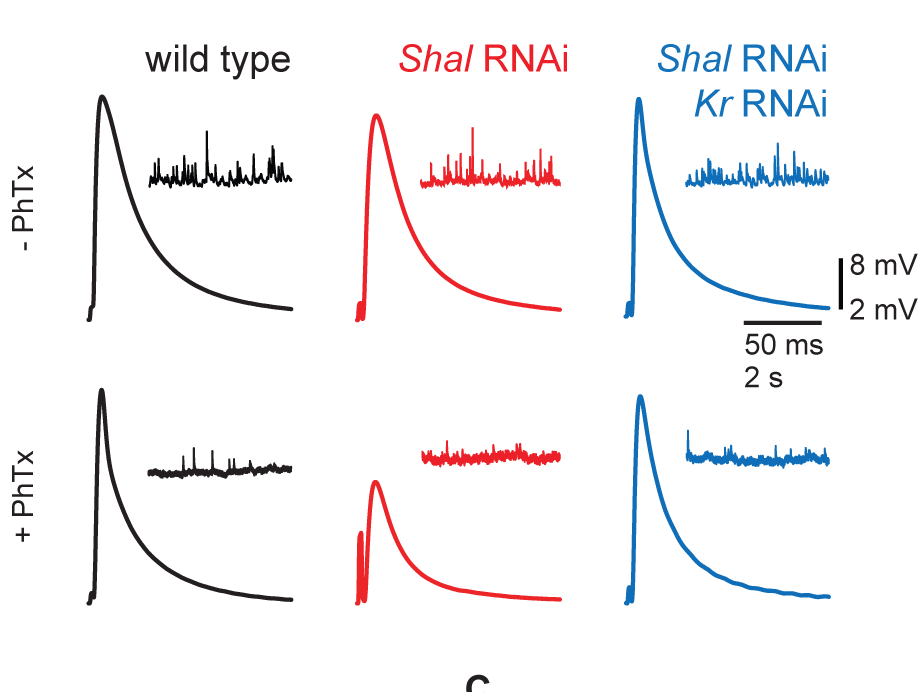
Krüppel mediates the selective rebalancing of ion channel expression
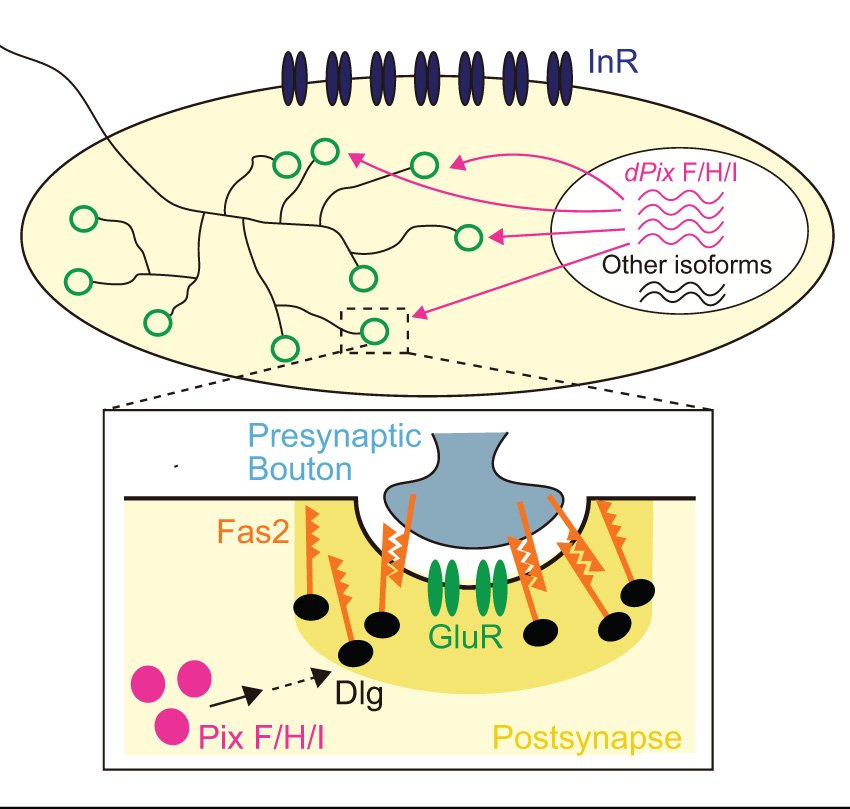
Parrish, J.Z.*†, Kim, C.K.*†, Tang, L., Bergquist, S., Wang, T., DeRisi, J.L., Jan, L.Y., Jan, Y.N., and Davis, G.W† (2014). Neuron 82: 537-544. PMCID: PMC4104505. *authors contributed equally, †authors for correspondence
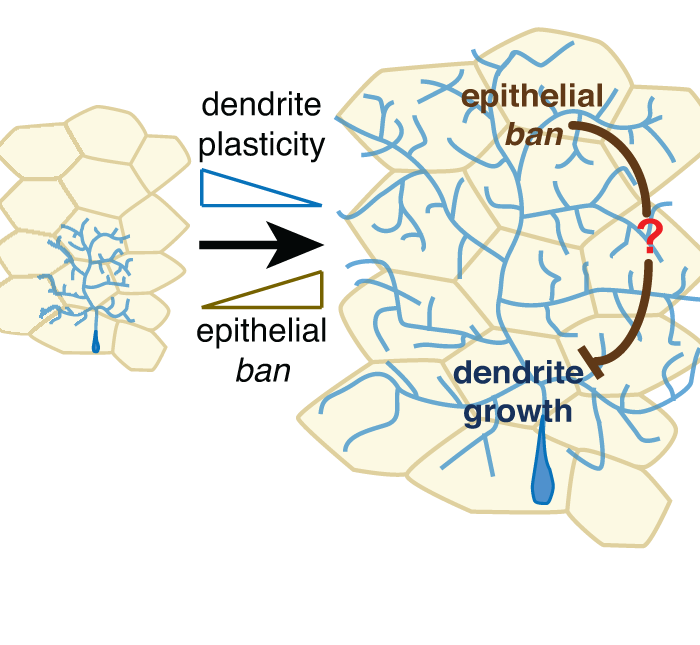
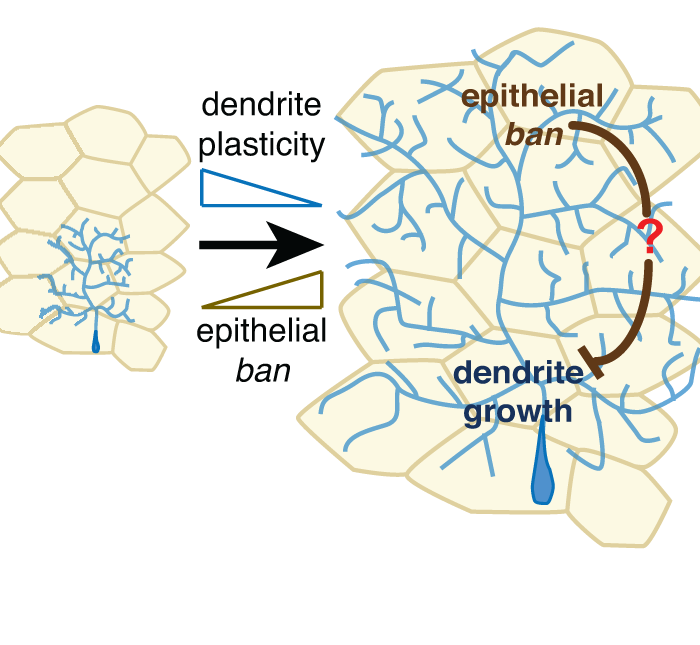
The microRNA bantam regulates a developmental transition in epithelial cells that restricts sensory dendrite growth
Jiang, N., Soba, P., Parker, E., Kim, C.K., and Parrish, J.Z. (2014). Development 141:2657-68. PMCID: PMC4067962. (see also: Keeping dendrites in check, Development 2014 141:e1302 )
See also Development editorial highlight: Keeping dendrites in check
2011
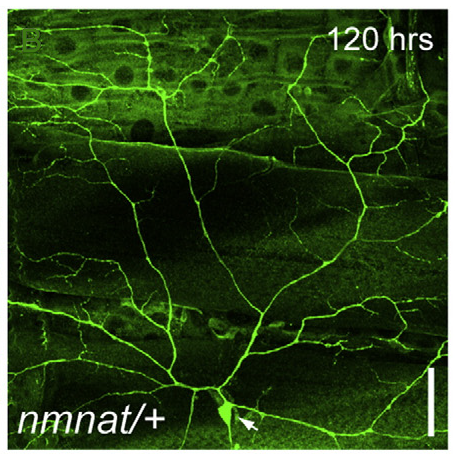
Nmnat exerts neuroprotective effects in axons and dendrites
Yuhui Wen, Jay Z. Parrish, Ruina He, R. Grace Zhai, and Michael D. Kim. (2011) Mol Cel Neurosci. 48:1-8
http://www.ncbi.nlm.nih.gov/pmc/articles/PMC3152617/
Jan Lab, Univ. of California San Francisco (2003-2009)
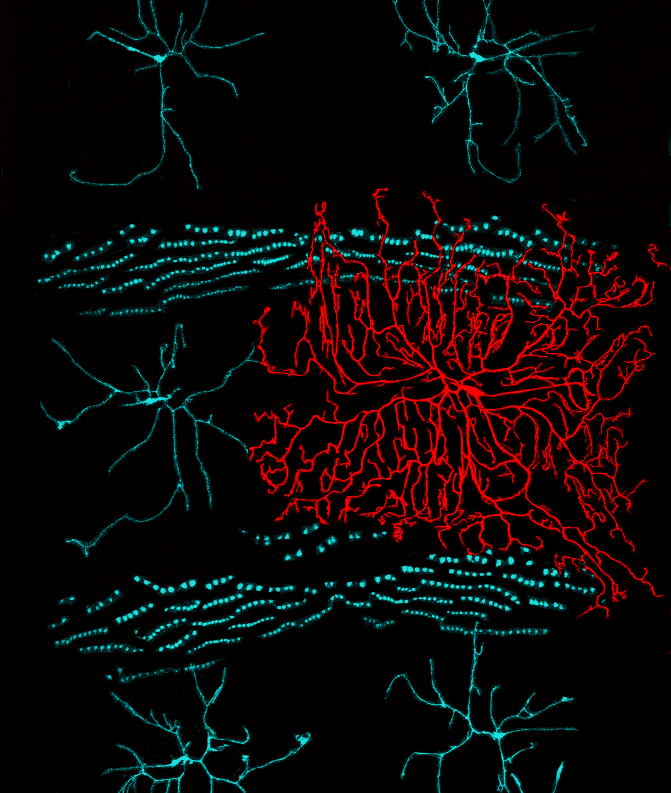
The microRNA bantam functions in epithelial cells to regulate scaling growth of dendrite arbors in Drosophila sensory neurons
Parrish JZ, Xu P, Kim CC, Jan LY, Jan YN (2009) Neuron 63:788-802
See also Nature Reviews Neuroscience research highlight: Scaling with microRNAs
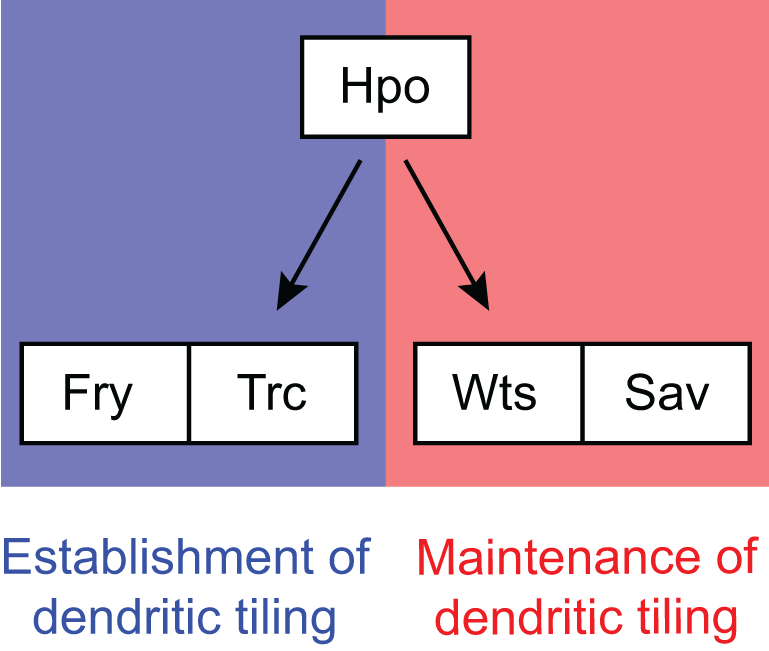
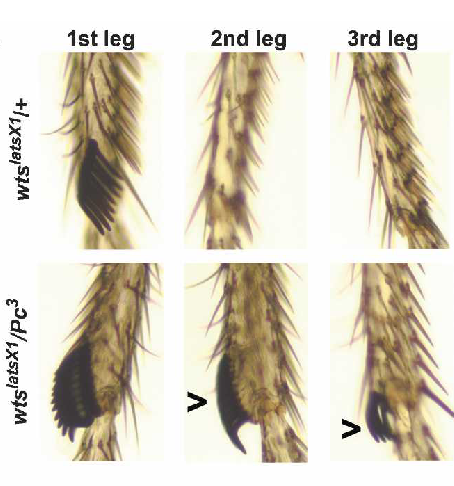
Polycomb genes interact with the tumor suppressor genes hippo and warts in the maintenance of Drosophila sensory dendrites
Parrish JZ, Emoto K, Jan LY, Jan YN. (2007) Genes and Development 21:956-972
http://www.ncbi.nlm.nih.gov/pubmed/17437999
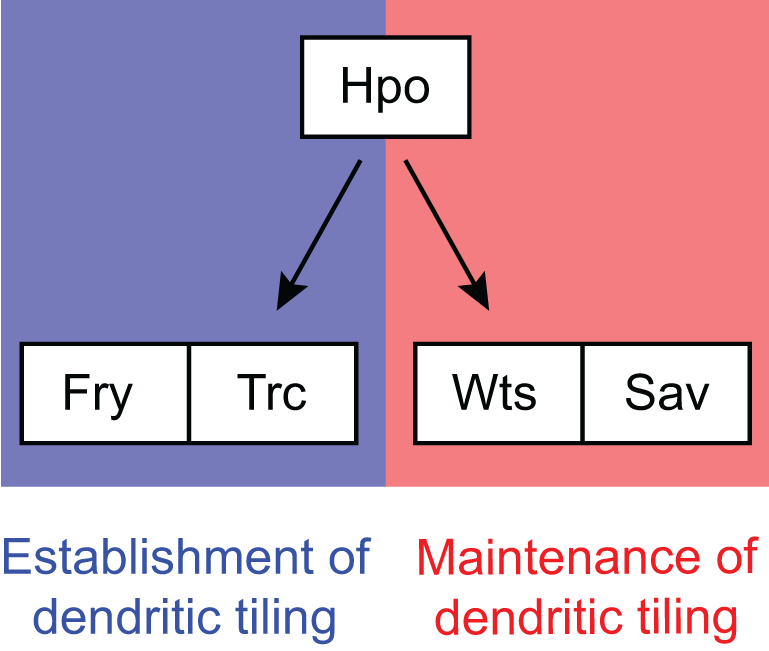
The tumour suppressor Hippo acts with the NDR kinases in dendritic tiling and maintenance
Emoto K, Parrish JZ, Jan LY, Jan YN (2006) Nature 443:210-213
http://www.ncbi.nlm.nih.gov/pubmed/16906135
Genome-wide analyses identify transcription factors required for proper morphogenesis of Drosophila sensory neuron dendrites.
Parrish JZ, Kim, MD, Jan LY, Jan YN (2006) Genes & Development 20:820-835
See also Cell Leading Edge and Genome Biology minireview: Transcriptional control of dendritic patterning in Drosophila neurons